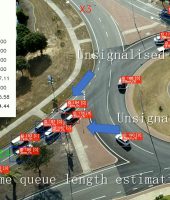
Charting how we feed a future world
Climate change is a threat to national and global food security, with generally decreased agricultural water supplies and rising temperatures affecting major agricultural yields including staple crops like canola or bread wheat. This has serious implications for Australia’s own food supply as well as its export market. Australian canola exports are forecast to increase by 42 per cent in 2020-2021, which would represent AU$1.35 billion to the Australian economy.
To help transform crop practices and develop more resilient crop varieties, we need to better understand the genetic make-up and breeding capabilities of these crops. Working under Professor David Edwards, Philipp Bayer and a team at the University of Western Australia are using genome sequencing to speed up the time it takes to develop, test, and cultivate new crops to feed us into the future.
Challenge
Philipp Bayer and the Applied Bioinformatics team under Professor David Edwards at the University of Western Australia are tackling the challenge of how to feed a future world, using next-generation genomic sequencing technology and bioinformatics.
This research involves coming up with ways to breed plants in our changing climate and develop new plants that are less prone to problems caused by a changing climate. But conventional plant breeding and cultivation processes — something that has historically taken decades — will not be enough to help feed a growing global population. The Applied Bioinformatics team is aiming to make it possible to develop new plant cultivars within a matter of years.
The team focuses on large plant genomes such as wheat, brassica, and chickpea. As part of this work, the Applied Bioinformatics team develops custom algorithms and analysis pipelines in order to gain a greater understanding of the complex plant genomes and their expressions.
Looking at thousands or hundreds of thousands of individual plant genomes at the same time generates massive amounts of data and takes time to analyse. Each plant genome must be pulled apart and compared to the reference genome to determine how variants might transform the gene.
Solution
Supercomputing is critical to Philipp’s research because of the scale of data he analyses and the complexity of the work.
“We’re working with humongous amounts of data — in the hundreds of terabytes of data. Just five years ago, we wouldn’t have been able to do the work we’re doing today. Supercomputers and this scale of computational power are essential to things like our genome assembly and pangenomics work,” Bayer says.
The team recently finished a project with 1,000 genomes which generated a few hundred terabytes of data. In collaboration with Professor Rajeev Varshney, ICRISAT in India, the team is looking to expand its work even further through a dataset of 30,000 chickpeas, which is going to generate approximately one petabyte of data.
The Applied Bioinformatics team is also deploying machine learning and deep learning techniques to further the impact of their research. Using Pawsey’s Topaz GPU cluster, Bayer is able to deploy machine learning algorithms that link genetic variants with traits of interest to farmers and breeders.
“We’re working with industry partners to do genomic prediction using deep learning. This involves five million data points per individual and 100 times over — then running machine learning techniques on that data. We run this on Topaz since the GPU compute power allows us to run state-of-the-art deep learning algorithms to achieve the highest prediction accuracy possible.”
Outcome
The scale and complexity of Bayer’s work have only been made possible by using supercomputers, bringing computations that are normally impossible on a laptop computer down to a few hours of work. This has expanded what his team is able to undertake and opened the door to bigger and more sophisticated projects.
“We’ve been able to scale our research to the next level. It’s now possible for us to look at the genomes of hundreds of thousands of individuals at the same time, which means we can map all the genetic variations of a whole population in just a few weeks.”
Project Leader.
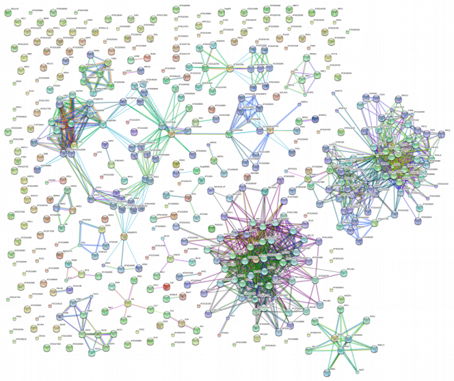
Gene clusters - sourced by Prof Edwards
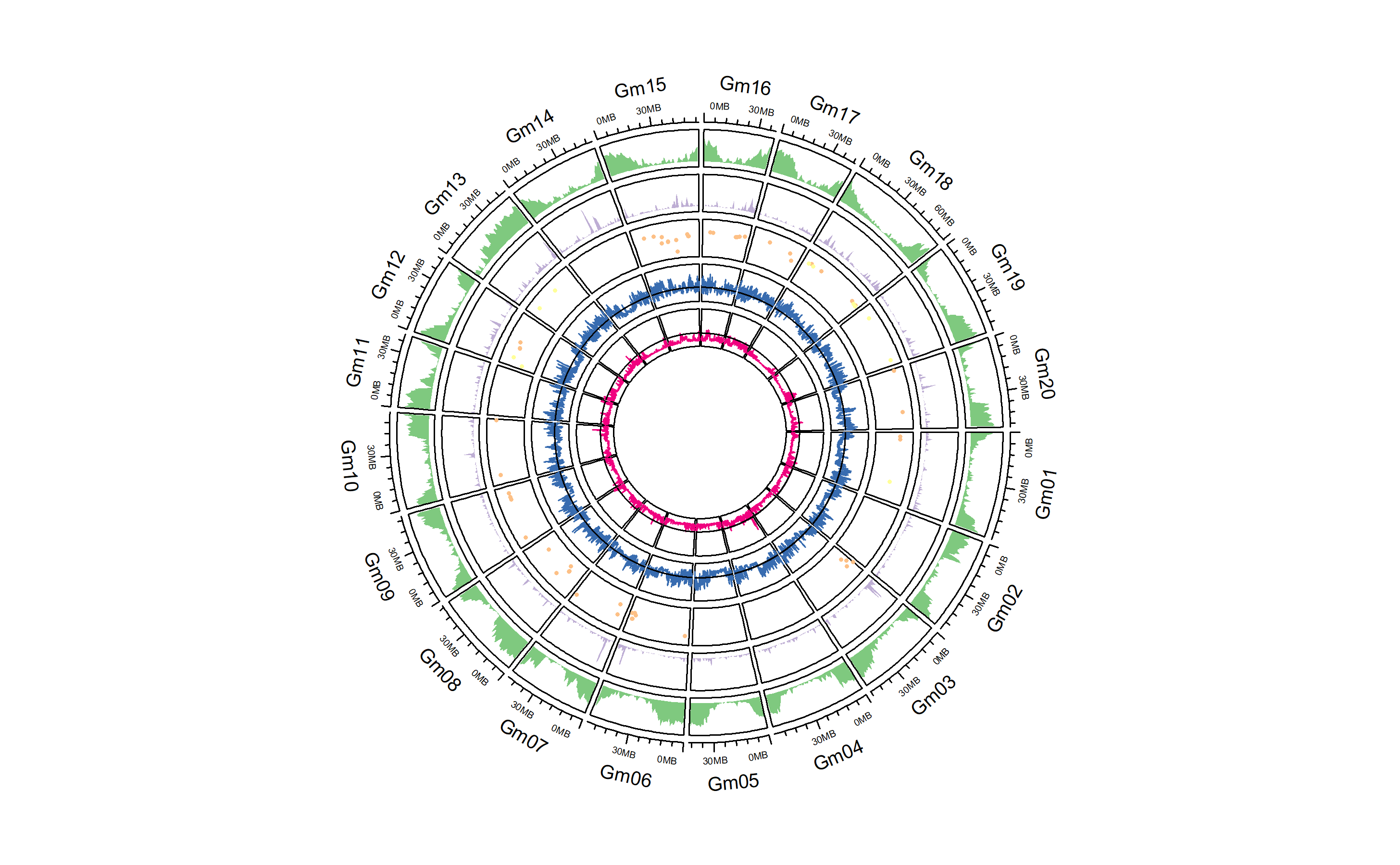
a Circos plot of the soybean genome showing various measurements of selective pressure