Clostridium difficile
This project seeks to develop a comprehensive understanding of the molecular epidemiology and genetic factors, including antimicrobial resistance (AMR), driving the evolution and transmission of the enteric pathogen Clostridium difficile in Australia.
Area of science
Microbiology
Systems used
Nimbus
Applications used
BioLinux (microbial genomics tools and bioinformatics)The Challenge
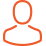
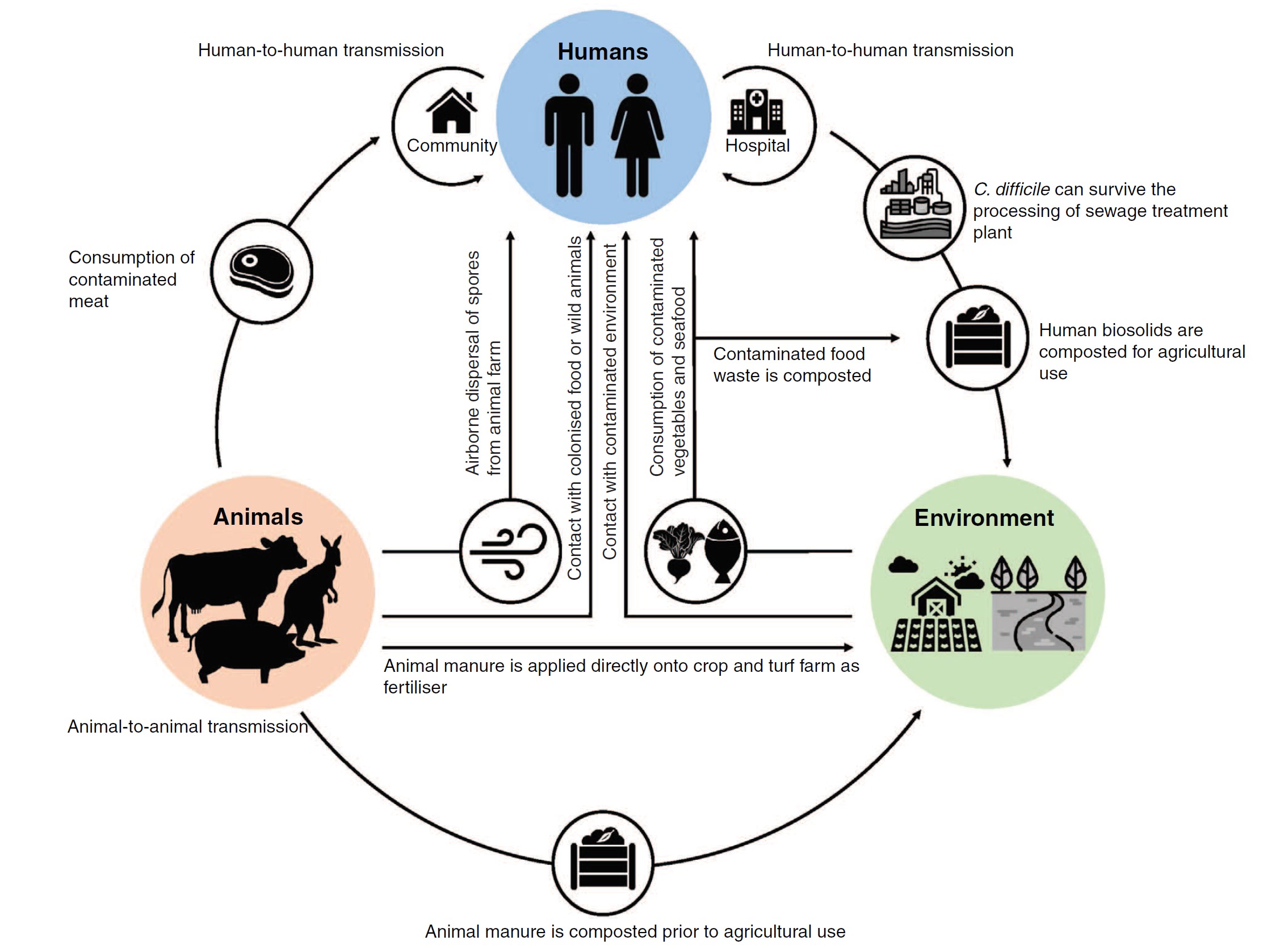
My research focus is the enteric bacterium Clostridium difficile – a formidable pathogen and the leading cause of life-threatening infectious diarrhoea in humans and many production animal species. The US Centers for Disease Control and Prevention (CDC) ranks C. difficile as the most important antimicrobial resistant threat to public health in the USA with 250,000 infections and 14,000 deaths per year, and annual excess medical costs totalling $1 billion. Clostridium difficile infection (CDI) has changed dramatically over the last 15 years and CDI may now have a food-borne aetiology with concomitant zoonotic transmission.
The Solution
My current research project utilises advanced genomics and a world-leading C. difficile strain collection to advance understanding of the epidemiological and genetic factors such as antimicrobial resistance (AMR) driving the evolution and transmission of C. difficile. Nimbus provides the computing power to conduct state-of-the-art genomic-based investigations into the evolutionary history and genetic diversity of C. difficile isolates originating from humans, animals, and their shared environment in Australia. Specifically, Nimbus will be used to run high-throughput algorithms for de novo assembly of bacterial genomes, multiple sequence alignment, read mapping and variant analysis, pan-genome analysis as well as very large Bayesian and maximum-likelihood based phylogenetic analyses.
The Outcome
Using the computing and cloud resources at the Pawsey I have optimised a pipeline for high-resolution core genome single nucleotide variant (SNV) analysis of C. difficile isolated from humans, animals and their shared environment. SNV analysis is highly discriminatory and the gold-standard method for reconstructing the fine-scale transmission networks and evolutionary history of bacterial pathogens. Applying this approach to a single critically important lineage of C. difficile that causes disease in humans and pigs in Australia (RT014), I have shown that these populations share a recent evolutionary history with the first evidence of interspecies transmission of C. difficile between pigs and humans in Australia. This information will be crucial in guiding the development of effective infection prevention and control strategies, and public health interventions, measures which will reduce the incidence and mortality of C. difficile infection (CDI) in humans and animals, thus enhancing Australia’s biosecurity.
List of Publications from this project
Putsathit P, Neela VK, Joseph NMS, Oiu PT, Ngamwongsatit B, Knight DR, Riley TV. 2019. Molecular epidemiology of Clostridium difficile isolated from piglets. Vet Microbiol 237:108408.
Androga GO, Knight DR, Hutton ML, Mileto SJ, James ML, Evans C, Lyras D, Chang BJ, Foster NF, Riley TV. 2019. In silico, in vitro and in vivo analysis of putative virulence factors identified in large clostridial toxin-negative, binary toxin-producing C. difficile strains. Anaerobe 60:102083.
Hong S, Knight DR, Chang B, Carman RJ, Riley TV. 2019. Phenotypic characterisation of C. difficile PCR ribotype 251, an emerging multi-locus sequence type clade 2 strain in Australia. Anaerobe 60:102066.
Imwattana K, Knight DR, Kullin B, Collins DA, Putsathit P, Kiratisin P, Riley TV. 2019. Clostridium difficile ribotype 017 – characterization, evolution and epidemiology of the dominant strain in Asia. Emerg Microbes Infect 8:796-807.
Azimirad M, Gholami F, Yadegar A, Knight DR, Shamloei S, Aghdaei HA, Zali MR. 2019. Prevalence and characterization of Clostridium perfringens toxinotypes among patients with antibiotic-associated diarrhea in Iran. Sci Rep 9:7792.
Knight DR, Kullin B, Androga GO, Barbut F, Eckert C, Johnson S, Spigaglia P, Tateda K, Tsai PJ, Riley TV. 2019. Evolutionary and genomic insights into C. difficile sequence type 11: a diverse, zoonotic and antimicrobial-resistant lineage of global One Health importance. mBio 10:e00446-19.
Wehrhahn MC, Keighley C, Kurtovic J, Knight DR, Hong S, Hutton M, Lyras D, Wang Q, Leong R, Borody T, Edye M, Riley TV. 2019. A series of three cases of severe Clostridium difficile infection in Australia associated with a binary toxin producing clade 2 ribotype 251 strain. Anaerobe 55:119-123.
Androga GO, Knight DR†, Lim SC, Foster NF, Riley TV. 2018. Antimicrobial resistance in large clostridial toxin-negative, binary toxin-positive Clostridium difficile ribotypes. Anaerobe 54:55-60.
Roshan N, Riley TV, Knight DR, Steer JH, Hammer KA. 2019. Natural products show diverse mechanisms of action against C. difficile. J. Appl. Microbiol. 126:468-479.
Amy J, Bulach D, Knight DR, Riley TV, Johanesen P, Lyras D. 2018. Identification of large cryptic plasmids in Clostridioides (Clostridium) difficile. Plasmid 96-97:25-38.
McGovern AM, Androga GO, Knight DR, Watson MW, Elliott B, Foster NF, Chang BJ, Riley TV. 2017. Prevalence of binary toxin positive Clostridium difficile in diarrhoeal humans in the absence of epidemic ribotype 027. PLoS One 12:e0187658.
Knight DR, Squire MM, Collins DA, Riley TV. 2017. Genome analysis of Clostridium difficile PCR ribotype 014 lineage in Australian pigs and humans reveals a diverse genetic repertoire and signatures of long-range interspecies transmission. Front Microbiol 7:2138.
McGovern AM, Foster NF, Pereira AL, Knight DR, Elliott B, Chang BJ, Riley TV. 2016. Human C. difficile infection caused by a livestock-associated ribotype 237 strain in Western Australia. JMM Case Rep 3.
Knight DR, Androga GA, Ballard SA, Howden BP, Riley TV. 2016. A phenotypically silent vanB2 operon carried on a Tn1549-like element in Clostridium difficile. mSphere 1:e00177-16.
Knight DR, Hart J, Gottardo NG, Eyre DW, Crook DW, Riley TV. 2015. Two cases of C. difficile infection in unrelated oncology patients attributable to a single clone of C. difficile ribotype 126. JMM Case Rep 2.
Knight DR, Elliott B, Chang B, Perkins TT, Riley TV. 2015. Diversity and evolution in the genome of Clostridium difficile. Clin Microbiol Rev 28:721-741.