Investigating Membrane Protein Dynamics, Regulation and Substrate Recognition
The overall aim of our research is to understand the molecular role of the cell membrane in regulating cellular processes, and the biochemical alterations that lead to disease. The fundamental aim of this work is to characterise lipid, protein and small molecule interactions in physiologically relevant membrane systems to further our understanding of normal cellular function, and as potential targets for therapeutic intervention. Further applications are the development of biologically inspired and bio-compatible materials.
Area of science
Biological Sciences, Chemical Biology, Chemical Sciences, Geosciences
Systems used
Magnus
Applications used
Gromacs 2016.1, Gromacs 2019.4 (these were both custom installed) Gromacs 2018 (this was as module on Zeus)The Challenge
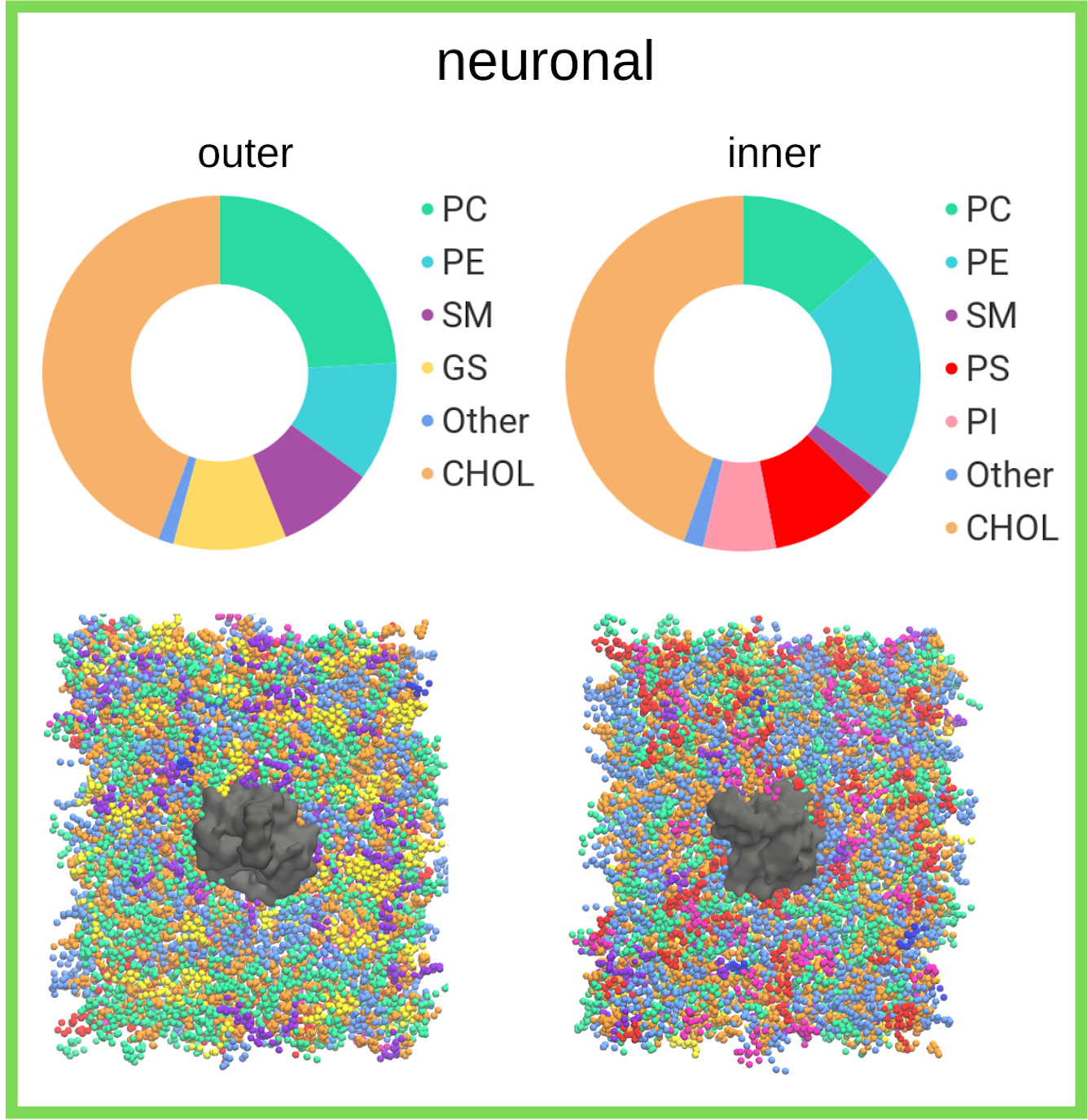
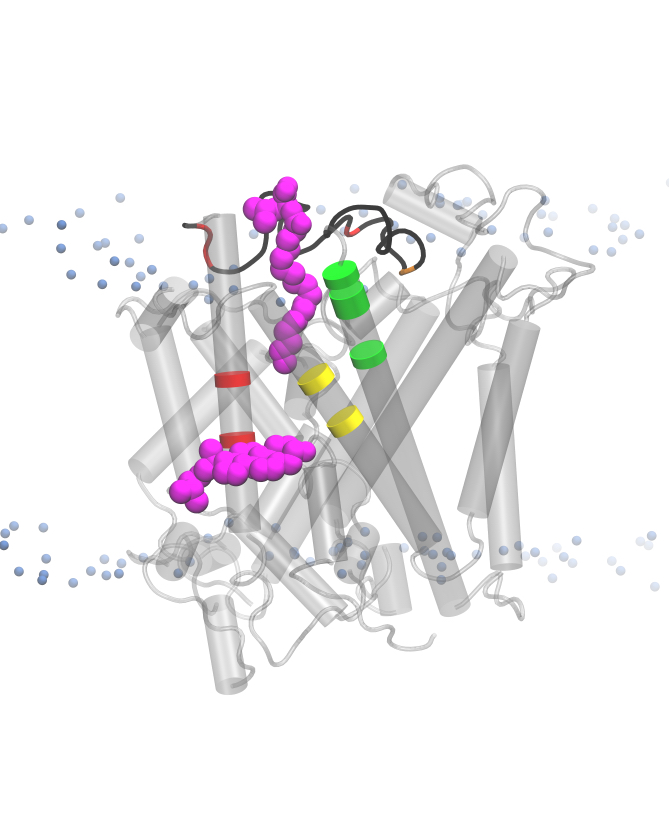
The research undertaken was focused primarily on understanding the local chemical environment of the neurotransmitter transporter, GlyT2, which is involved in the pain pathway. This research investigated lipid self-organisation within a complex model of the neuronal membrane and characterised the differential association of lipid species around GlyT2 and other neurotransmitter transporters. The research worked to develop a structure function activity relationship for novel lipid-based inhibitors of GlyT2, which will aid in the development of new analgesics to treat chronic pain. These inhibitors vary in the structure of the lipid tail and these changes in the lipid tail have been shown experimentally to lead to inactivation of the lipid inhibitor.
Further extensions of this work involved the use of lipids and micelles in enzyme mimicry
The Solution
Current experimental techniques cannot resolve both temporal and spatial changes at atomic resolution. Molecular dynamics (MD) simulations are currently the only technique that is able to characterize the time evolution of these systems to predict how changes in the chemical environment (e.g., presence of ligands) drives conformational dynamics. The resulting trajectories will be combined with structural and biochemical data to characterise the molecular interactions that govern function
The Outcome
This research provides unprecedented detail on the local remodeling of the neuronal membrane in the area surrounding neurotransmitter transporters such as GlyT2. It provides details on the binding interactions of lipid-based inhibitors to GlyT2 that can be further examined through experimental techniques. Finally, characterization of self-assembling interactions in lipid environments gives insights into the interactions governing chemical specificity which can be exploited for the design of biologically inspired materials.
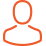
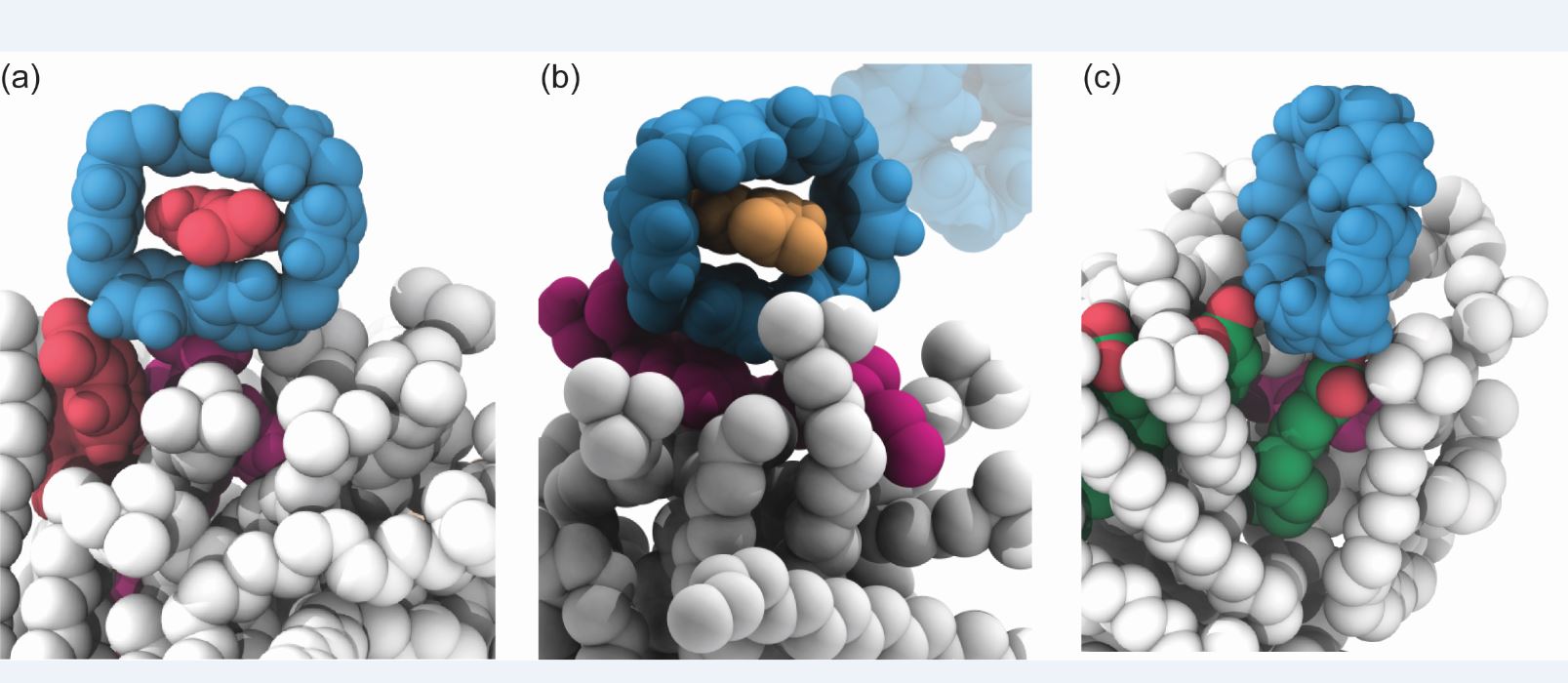
 Enzyme mimicry in a lipidic environment. Lipid micellar systems containing electron rich aromatics such as a) benzoate or b) nitrophenolate anions bind to cyclobis(paraquat-p-phenylene) (bluebox) to undergo selective hydrolysis. In contrast, electron rich alykl products such as c) dodecanoate anions do not bind to bluebox, preventing catalysis.
Enzyme mimicry in a lipidic environment. Lipid micellar systems containing electron rich aromatics such as a) benzoate or b) nitrophenolate anions bind to cyclobis(paraquat-p-phenylene) (bluebox) to undergo selective hydrolysis. In contrast, electron rich alykl products such as c) dodecanoate anions do not bind to bluebox, preventing catalysis.
List of Publications
1) Shannon N Mostyn, Katie A Wilson, Alexandra Schumann-Gillett, Zachary J Frangos, Susan Shimmon, Tristan Rawling, Renae M Ryan, Megan L O’Mara, Robert J Vandenberg. Identification of an allosteric binding site on the human glycine transporter, GlyT2, for bioactive lipid analgesics. eLife. doi:10.7554/eLife.47150
2) Mitchell D Nothling, Zeyun Xiao, Nicholas S Hill, Mitchell T Blyth, Ayana Bhaskaran, Marc-Antoine Sani, Andrea Espinosa-Gomez, Kevin Ngov, Jonathan White, Tim Buscher, Frances Separovic, Megan L O’Mara, Michelle L Coote, Luke A Connal. A multifunctional surfactant catalyst inspired by hydrolases. Science Advances 6:eaaz040