Accelerating the Design of Novel Catalysts and Drugs through Computational Chemistry
Presently, the search for improved materials, catalysts or drug molecules has relied almost exclusively on experiment that is costly both in terms of manpower and resources. Our research interest lies in the development and application of state-of-the-art computational methods to help tackle these challenges. Topics of particular interest include the development and application of cost-effective approaches to modelling supramolecular chemistry in catalysis and medicine, reactivity on surfaces, biomolecular dynamics and allostery. Ultimately, our goal is to use computational chemistry to elucidate structure activity trends, and understand the origin of these relationships that will facilitate the design of better chemical reagents and drug molecules.
Area of science
Chemical Sciences, Chemistry, Computational Chemistry
Systems used
Magnus
Applications used
NAMD, ORCA and NWCHEMThe Challenge
Current computational methods lack predictive power for computation-driven design and discovery of drug molecules and chemical reagents.
The Solution
We are developing robust computational methods and efficient approximations to these methods that will address the deficiencies in existing models.
The Outcome
The computational resources at Pawsey Supercomputing Centre will enable us to generate highly accurate quantum chemistry data to test and validate our computational approaches across a broad range of systems. It will also provide the platform for implementing our methods.
List of Publications
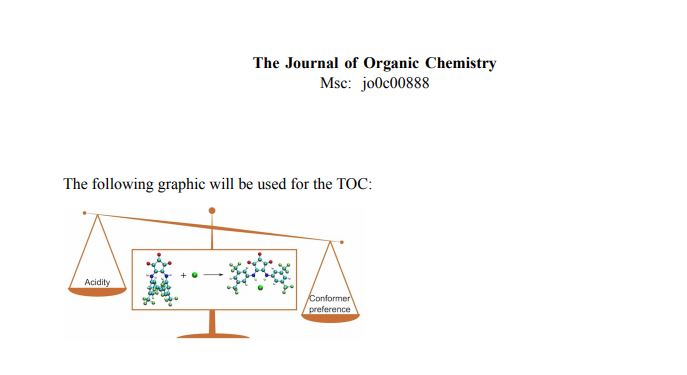
(1) I. Sandler, F. A. Larik, N. Mallo, J. E. Beves, J. Ho.
Anion Binding Affinity – Acidity versus Conformational Effects.
J. Org. Chem. (in press) DOI: 10.1021/acs.joc.0c00888
(2) J. Chen., B. Chan, Y. Shao, J. Ho
How Accurate are Approximate Quantum Chemical Methods at Modelling Solute-Solvent Interactions in Solvated Clusters?
Phys. Chem. Chem. Phys. 2020, 22, 3855.
(3) V. Kundi, J. Ho
Predicting Octanol−Water Partition Coefficients: Are Quantum Mechanical Implicit Solvent Models Better than Empirical Fragment- Based Methods?
J. Phys. Chem. B, 2019, 123, 6810.
(4) J. Chen., Y. Shao, J. Ho
Are Explicit Solvent Models More Accurate than Implicit Solvent Models? A Case Study on the Menschutkin Reaction.
J. Phys. Chem. A 2019, 123, 5580
(5) S. Wong et al.
Just add Sugar for Carbohydrate Induced Self-Assembly of Curcumin.
Nature Comm. 2019, 10, 1.
(6) H. Wang, M. G. Leeming, J. Ho, W. A. Donald
Origin and PredictionofHighlySpecific Bond Cleavage Sites inthe ThermalActivation of Intact Protein Ions
Chem. Eur. J. 2019, 25, 823