Ascochyta-rabiae
Ascochyta blight caused by the fungus Ascochyta rabiei is a major foliar disease affecting chickpea farmers in Australia and worldwide
Area of science
Plant biology
Systems used
Magnus, Zeus and Nimbus
Applications used
canu; pilon; trinity; hisat2; trimommatic; fastqc; R; inteproscanThe Challenge
Australian A. rabiei population has low genotypic diversity evinced by the presence of only one mating type. However, isolates have become more diverse in aggressiveness, affecting even the newly released resistant chickpea varieties.
Our aim was to employ genomics techniques to better understand the mechanisms adopted by A. rabiei to evolve increased pathogenicity and to identify genomic factors that potentially support the heterogeneity in aggressiveness and pathogenicity.
The Solution
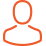
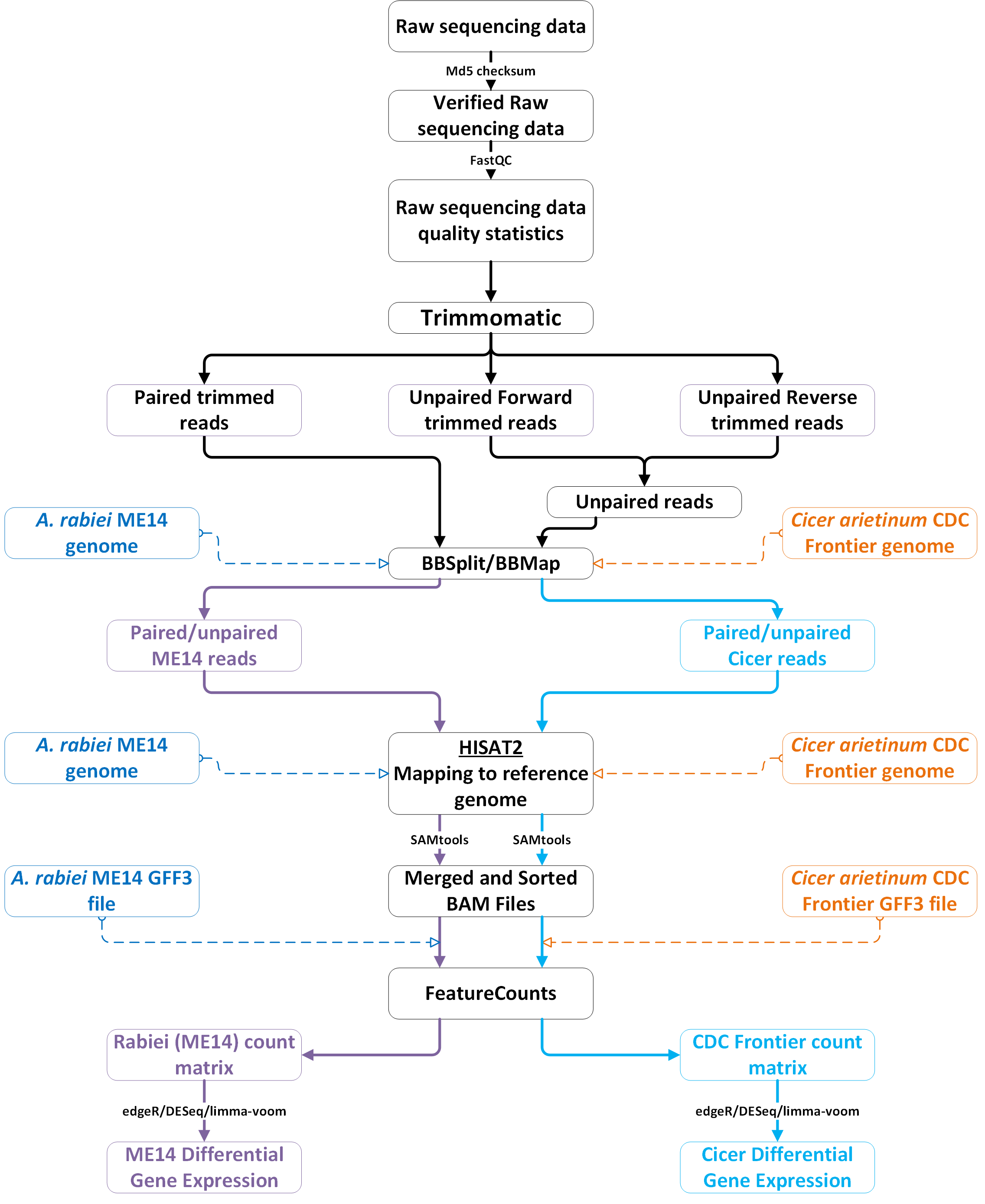
We assembled near-complete genomes using high resolution long-read whole-genome sequencing with short-read polishing and performed in-depth comparative analyses of Australian isolates exhibiting diverse aggressiveness. We also performed RNAseq analysis to identify genes that are differentially expressed in A. rabiei during the interaction with chickpea.
The Outcome
Parallel comparison between relatively large genomes is a computing-intensive task that could not be possible on a single PC. With the extra computing capacity at Pawsey Centre, we were able to assemble and compare whole genomes and transcriptomes of A. rabiei, leading to the identification of genes and genomic hotspots, including polymorphisms, that could be attributed to increased pathogenicity among isolates.

Figure 1. The completeness and quality of A. rabiei draft genomes. A. BUSCO summary of the genome scaffolds. B. QUAST Nx plot showing the changes in Nx metrics (bp) relative to the genome proportion (%) for the different assemblies. C. Percentage GC-content distribution across the different assemblies as calculated by OcculterCut